
Scientometrics and Network Science
Currently, under the guidance of Prof. George Chacko and Prof. Tandy Warnow, I am working with large networks of research papers linked through citation. The largest network I have worked with was a citation network of ~75 million nodes and ~1 billion edges (OpenCitations). The specific projects I'm working on are below:
- High-performance well-connected community detection (link)
- Using multi-dimensional metrics to analyze publication impact both at the community and network scale
- Simulating citation network growth and encoding citation behavior.
.
Image: A rat made out of citations. Each node is a paper, and each edge is a citation. A lab rat who is merely trying to understand its own behavior.
Image generated by DALL-E
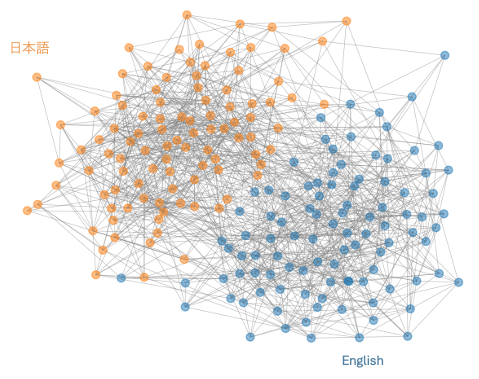
Understanding Community Formation in Science through Cross-lingual Citation
This project was worked on mainly during my time at the National Institute of Informatics (国立情報学研究所) in Tokyo, under the guidance of Prof. Chifumi Nishioka.
In this project, we explored how cross lingual citation can divide a research citation into communities. Moreover, we explored how messenger nodes across languages can bridge communities. We used a dataset of 7 million papers from a corroborated dataset across OpenCitations, CrossRef, and OpenAlex.
Image: A visualization of a citation network across languages. Each color represents a different language, and the size of the node represents the number of citations the paper has. NOTE: This is a mockup and not the actual visualization.
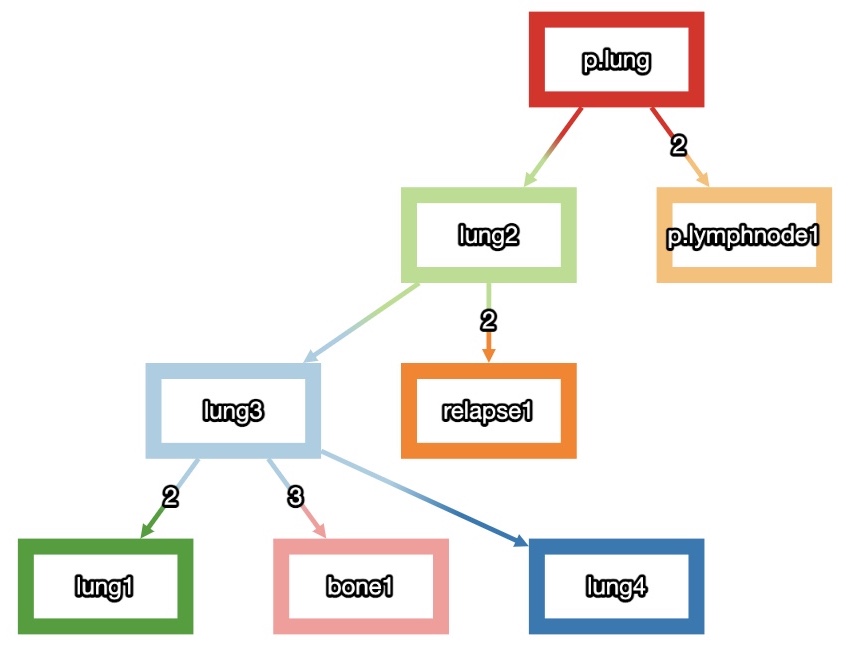
Migration History Inference of Metastatic Cancers
I have also been working with Prof. Mohammed El-Kebir on using clonal trees inferred from variant allele frequencies (VAFs) to infer anatomic labelings of nodes in the clonal tree and ultimately a migration graph. Since his paper on MACHINA in 2018, we have expanded the solution space to return multiple possible migration histories per primary tumor location.
I created this visualizer. More than simply providing visual aid for the algorithm, it allows users (researchers, oncologists, etc.) to filter through the solution space and enforce priors such as known migrations and/or absences of migrations.
Image: Migration history inferred for a patient recorded in the TracerX consortium.
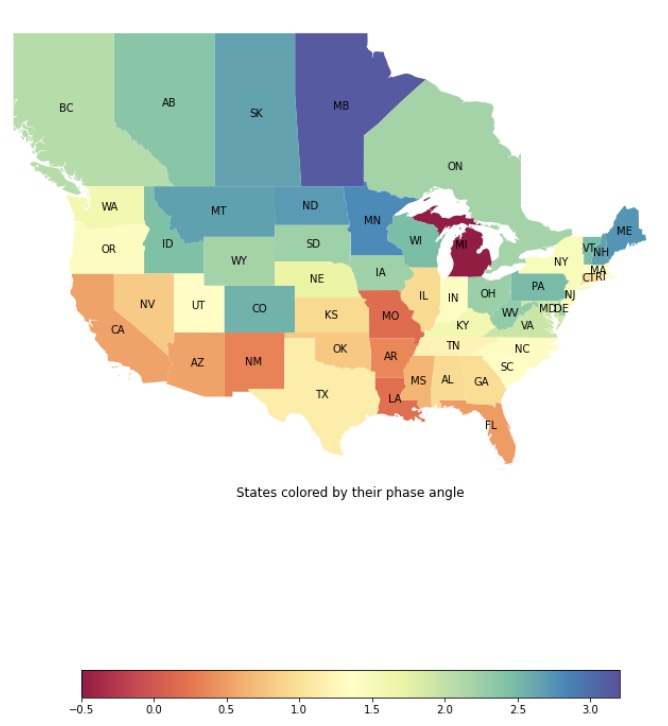
Cyclicity Analysis on COVID in North America
Cyclicity analysis is the technique of aggregating regional linear time series to map spread of a signal over a medium. Cyclicity analysis was traditionally used in neuroscience to infer spread of a signal across the brain given individual synaptic potentials.
However, we now use Cyclicity Analysis to understand the spread of different variants of COVID. Using American and Canadian provincial COVID case time series, spread is mapped across North America.
Image: Inferred spread of the Delta variant of COVID. Redder colors indicate earlier states in the wave, and bluer colors indicate later states in the wave.
Previous Research Experiences
(2020-2022) Development of an Instructional Alexa Skill for Older Adults [Department of Community Health, University of Illinois at Urbana-Champaign]
(2017) A Smart Power Outlet That Benefits From Real-Time Pricing [Grainger College of Engineering, University of Illinois at Urbana-Champaign]
Papers
Publications
- Well-Connected Communities in Real-World Networks
Minhyuk Park*, Yasamin Tabatabaee*, Vikram Ramavarapu*, Baqiao Liu, Vidya Kamath Pailodi, Rajiv Ramachandran, Dmitriy Korobskiy, Fabio Ayres, George Chacko, Tandy Warnow (2024)
Proceedings in COMPLEX Networks 2023. Accepted at PLoS (Link) - CM++ - A Meta-method for Well-Connected Community Detection
Vikram Ramavarapu, Fábio Jose Ayres, Minhyuk Park, Vidya Kamath Pailodi, João Alfredo Cardoso Lamy, Tandy Warnow, and George Chacko (2023)
In submission at the Journal of Open Source Software (JOSS) (Link) - A Smart Power Outlet for Electric Devices that can Benefit from Real-Time Pricing
V.P. Ramavarapu, R. Sreenivas, R. Sowers (2017)
Initially a conference article in The Proceedings of the 2017 International Conference on Control, Electronics, Renewable Energy and Communications (ICCEREC). Published at IEEE (Link)
Presentations
-
Exploration of Multi-Lingual Community Structure in Scholarly Articles
Vikram Ramavarapu, Chifumi Nishioka (2024)
To be presented at the ACM/IEEE Joint Conference on Digital Libraries (JCDL) 2024 -
Inferring Migration Histories of Metastatic Cancers
Mohammed El-Kebir, Mrinmoy Roddur, Vikram Ramavarapu
Presented at the Banff International Research Station (BIRS) 2023